Making QBIC Faster
(viewing version)
Making QBIC Faster
Efficient Content-based Image Retrieval
John Shepherd
joint work with
Nimrod Megiddo, Xiaoming Zhu
IBM Almaden Research Center (Feb-Aug,1998)
Overview
- QBIC and Content-based Image Retrieval
- High-dimensional k Nearest Neighbours Problem
- Database Indexing Schemes and HDkNN Retrieval
- The Curveix Approach ... Preliminary Results
- Conclusions and Future Work
The QBIC System
QBIC was developed around 1983 as an experiment in image retrieval.
The aim was to build a system for images that provide analogy
to text retrieval operation ``Find documents like this one''.
The focus was on developing measures of image similarity that:
- corresponded somewhat to human perception of "similarity"
- were not too expensive to compute
... The QBIC System
![[Diagram:pic/qbic]](pic/qbic.gif)
Content-based Image Retrieval
User supplies a description of desired image via feature vectors.
System returns a ranked list of "matching" images from DB.
![[Diagram:pic/cbir-sys]](pic/cbir-sys.gif)
The Problem
The goal of the QBIC project is to develop a system that
- returns images that closely match the user's intent
- does this quickly, even for very large image collections
So far, the system has been somewhat successful on
the first point and has limited itself to databases
of < 10,000 images.
One efficiency measure already implemented: an N  N table of
pre-computed distances to speed up retrieval within database.
N table of
pre-computed distances to speed up retrieval within database.
High-dimensional Nearest Neighbours Search
The problem of finding k similar images is an instance of
the more general problem ...
Given:
- a set of data points in a d-dimensional space
- a specific query point q in that space
find the k data points in the set that are closest to q.
A solution to this problem would be useful in several areas:
- information retrieval, multimedia databases, machine learning, ...
The Problem (Re-stated)
Inputs
- a database of N objects
(obj1, obj2, ..., objN)
- representations of objects as d-dimensional vectors
(vobj1, vobj2, ..., vobjN)
(each vector may represent one or more features of the object)
- a query object q with associated d-dimensional vector (vq)
- a distance measure D(vi,vj) : [0..1)
(D=0 -> vi=vj)
Outputs
- a list of the k nearest objects in the database
[a1, a2, ... ak]
- ordered by distance to query object
D(va1,vq)
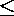 D(va2,vq)
D(va2,vq) 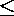 ...
... 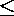 D(vak,vq)
D(vak,vq)
And all done as efficiently as possible ...
... where time is the most important efficiency factor
The Context
The solution has to work for ...
- number of dimensions d = 1 .. 500
- number of objects N = 104 .. 1010
- all values in query vector are specified (i.e. no partial-match)
- implementation runs on "standard" hardware/OS
(e.g. don't assume arrays of parallel disks, huge memory, ...)
- independent of distance measure (?)
- 100% retrieval accuracy (?)
And, most importantly, the solution should ...
- use minimal resources: i/o, cpu-time, space
- provide fast database updates
(important for some applications e.g. Magnifi)
The Costs
I/O cost
- the dominant cost (given current disk technology)
- measured in terms of pages read, not records read
CPU cost
- primarily related to computing distance measure D(vobj, vq)
- can be relatively expensive for some QBIC features
Space
- need to store data (feature vectors) within page-size chunks
- access methods typically require additional disk space
(which can be anywhere from 10% extra to >100% extra)
- intermediate data structures in memory?
Cost Parameters
We need to consider the following quantities:
| Name |
Meaning |
Typically |
| N |
number of objects in the database |
103 .. 1010 |
| d |
number of dimensions in object features |
1 .. 256 |
| P |
number of bytes per disk page |
512 .. 8K |
| V |
number of bytes for d-dimensional vector |
4d |
| O |
average number of bytes per stored object (key+vector) |
12+4d .. 256+4d |
NO |
average number of stored objects per page
 P/O P/O |
1 .. 512 |
| NP |
number of disk pages to hold stored objects
 N/NO N/NO |
50 .. 1010 |
| Topen |
time to open a database file |
10ms |
| TP |
time to read a page from disk into memory |
10ms |
| TD |
time to compute distance between two objects (using vectors) |
100us (?) |
| TK |
average time to find the page address for a given object |
10ms (?) |
The TK measure is related to the use of a DBMS
(e.g. dbm or DB2) to look up object vectors via the object
identifier/key (the file name of the image).
Analysing Costs
We (pessimistically) assume
that there is no effective caching of pages read from the database.
That is, accessing n object vectors (chosen at random) is
going to result in n page reads.
This gives an upper-bound on the actual cost.
The real cost will typically be less due to clustering and caching.
On the other hand, we also assume that if the access to
the vectors is via a sequential scan, then each page in
the database file will be read exactly once.
Measuring Costs
Unix has useful tools for measuring/analysing CPU costs (gprof).
Unix (AIX) is not helpful in measuring I/O costs
(the rusage structure doesn't seem to count block input/output)
We use a combination of
- our own I/O monitoring (to count
read, write ops)
- profiling (fine-grained analysis of cpu-time costs)
- wall-clock time (the "bottom line" for users)
Current Approach
Given a query vector vq, current QBIC method is (essentially):
results = []; maxD = infinity
for each obj in the database
{
dist = D(vobj,vq)
if (#results < k or dist < maxD)
{
insert (obj,dist) into results
// results is sorted with length <= k
maxD = largest dist in results
}
}
Cost = Topen + NPTP + NTD
Note: If q is an image from the database, we can use a
pre-computed distance table to make this much faster.
Specific Goals for a Solution
Any reasonable solution aims to:
- reduce the number of objects considered (find candidates)
- ... which, in turn, reduces i / o and D computations
- but without excessive overhead in determining candidates
Some Potential Solutions
Partition-based
- use auxiliary (spatial) data structure to identify candidates
- space-partitioning methods: Grid file, k-d-b-tree, quadtree
- data-partitioning methods: R-tree, X-tree, SS-tree, TV-tree, ...
Approximation-based
- use approximating data structure to identify candidates
- signature files: VA-files
- projections: space-filling curves
A new approach, the pyramid technique
[Berchtold et al.,]
has elements of both partitioning and projection.
... Some Potential Solutions
Other Optimisations
- reduce I/O by reducing size of vectors (compression)
- reduce I/O by placing "similar" records together (clustering)
- reduce I/O by remembering previous pages (caching)
- reduce cpu by making D computation faster (ColorHist)
Tree-based Methods
Each leaf in the tree corresponds to a subregion of the d-dimensional space
Non-leaf nodes correspond to a union of subregions in children
Partitioned space containing data points:
![[Diagram:pic/space]](pic/space.gif)
... Tree-based Methods
Tree that induced above partitioning:
![[Diagram:pic/tree]](pic/tree.gif)
Above has no overlap among subregions; some partitioning schemes
permit overlap.
Above uses hyper-rectangles for partitioning;
other shapes (e.g. spheres, convex polyhedra) have been used.
Detailed summary/comparison of tree methods can be found in
[White & Jain]
,
[Kurniawati et al.]
Insertion with Trees
Given: a feature vector vobj for a new object.
Insertion procedure:
traverse tree to find leaf node n
whose region contains vobj
if (enough space in n)
{
insert vobj into corresponding page
}
else
{
need to perform local rearrangement in tree
by splitting points between n and n's parent
and n's siblings according to split heuristic
}
Split heuristics include:
- minimise total volume of bounding regions
- minimise variance in volume of regions
- minimise overlap among regions
- .....
Query with Trees
Given: query vector vq, index tree.
Method involves:
- a search over the tree
- a "nearest neighbour (NN) region"
(related to maxD in the original search method).
Consider the following space and two query points q1 and q2.
![[Diagram:pic/qspace]](pic/qspace.gif)
... Query with Trees
Aim of search: reduce NN-region quickly as possible
(no need to consider nodes which do not intersect NN-region)
Variations on search methods in literature:
- overall search order
e.g. depth-first vs. breadth-first
- order to check nodes
e.g. distance from query point to node centroid
- pruning criteria
e.g. minimum distance exceeds
maxD
May not achieve 100% accuracy; depends on pruning strategy.
Why Trees Don't Work (for d > 10)
[Weber and Blott]
derive some results on average performance of kNN search:
- (degeneration)
the complexity of searching via a partitioning scheme tends
to O(N) for sufficiently large d
- (complexity)
for every partitioning scheme, there exists some d
such that all blocks are accessed if dimensionality exceeds d
- (performance)
most data- and space-partitioning schemes perform worse than
sequential scan by the time d=10
Signature-based Methods (VA-Files)
Signature methods abandon the idea of search complexity better than O(N).
Use a file of signatures in parallel with main data file,
one signature for each vector.
![[Diagram:pic/sigfile]](pic/sigfile.gif)
Signatures have properties:
- (much) more compact than vectors
- provide approximation to information in vectors
One example is the Vector Approximation file by
[Weber and Blott.]
VA-File Signatures
Vector (b1, b2, ..., bd)
has corresponding signature
him(b1)him(b2)...him(bd)
where him gives the top-order m bits
This essentially:
- partitions each dimension into 2m regions
- maps vector into "identifier" of region that contains it
![[Diagram:pic/va-partition]](pic/va-partition.gif)
For non-equi-width regions, use mapping array to determine
partition boundaries.
Insertion with VA-Files
Given: a feature vector vobj for a new object.
Insertion is extremely simple.
sig = signature(vobj)
append vobj to data file
append sig to signature file
Storage overhead determined by m (e.g. 3/32).
Query with VA-Files
Given: query vector vq.
results = []; maxD = infinity;
for each sig in signature file
{
vnear = closest point to vq in region[sig]
dist = D(vnear,vq)
if (#results < k or dist < maxD)
{
dist = D(vobj,vq)
if (#results < k or dist < maxD)
{
insert (obj,dist) into results
maxD = largest dist in results
}
}
}
Cost = 2Topen + TP(Ns + fN) + TD(N + fN)
where
- Ns is number of pages in signature file
- f is filtering factor (0.0005 under ideal conditions)
Note: achieves 100% accuracy.
Why VA-Files Don't Work (for QBIC)
In the experiments reported by
[Weber and Blott]
they use uniform data distributions and achieve 0.0005 filtering.
In experiments with QBIC colour histograms, which have extremely skew
distributions, we were unable to achieve better than 0.10 filtering
on average.
And this was after "tuning" partition boundaries to place approximately
equal numbers of points in each partition.
Before tuning, filtering factor was 0.70.
Conclusion VA-file performance is very sensitive to data distribution.
VA-files will also perform poorly if TD is large
(we compute D for every signature)
Projection-based Methods (C-curves)
Achieve efficiency by
- projecting data space onto reduced dimensions
- utilising existing access methods (e.g. B-trees) to find objects in projection
The projection typically loses information, and so these methods
may give an approximate answer.
Our method, CurveIx (Curve Index),
is an example of this approach.
Proposed Approach: CurveIx
Further develops a method initially proposed in
[Megiddo and Shaft]
(And possibly earlier? ... the idea of space-filling curves has been
around for a while)
The basic idea:
- project d-dimensional vector onto 1-d space-filling curve
- collect set of curve-neighbours (should contain some k-NN)
- repeat above for several curves with different paths through space
- union of neighbour sets from all curves gives candidates
- retrieve and compute distance only for candidates
![[Diagram:pic/c-curves]](pic/c-curves.gif)
We want candidate set to contain most/all of k-NN ...
- how many curves do we need?
- how many neighbours do we examine on each curve?
Curve Mapping Functions
Each Ci has the following properties:
- maps d-dimensional vectors onto points on a line
i.e. [0,1)d
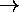 [0,1)
[0,1)
- mapping is one-to-one and defines an order on [0,1)d
- inverse mapping is continuous
Points within a small region of the curve are likely
to have been mapped from vectors that are close in d-space.
We generate multiple Cis by applying:
- permutations over vector elements
- translations of individual elements
Data Structures for CurveIx
| Ci |
a mapping function for each of the m
different space-filling curves (i=1..m)
|
| Db |
a database of N d-dimensional feature vectors;
each entry is a pair:
(imageId, vector)
imageId forms a primary key
|
| Index |
a database of Nm curve points; each point is represented
by a tuple
(curveId, point, imageId)
the pair (curveId, point) forms a primary key with an
ordering, where point = CcurveId(vimageId)
|
| vq |
feature vector for query q
|
Database Construction
For each image obj, insertion requires:
for i in 1..m
{
p = Ci(vobj)
insert (i, p, obj) into Index
}
insert (obj, vobj) into Db
Example: Search with 5 Curves
![[Diagram:pic/curves]](pic/curves.gif)
Finding k-NN (Simple Approach)
Given: Db , vq ,
C1 .. Cm , Index
for i in 1..m
{
p = Ci(vq)
lookup (i,p) in Index
fetch (i,p',j)
while p' "close to" p on curve i
{
collect next (i,p',j) as candidate
}
}
for each candidate j
{
lookup j in Db
fetch (j,vj)
d = D(vj , vq)
include j in k-NN if d small enough
}
Cost = 2Topen + mTsel + NfTP + NfTD
where
- Tsel is the cost of Index lookup using e.g. B-tree
- f is the filtering factor
CurveIx vs. Linear Scan
For linear scan:
Cost = Topen + NPTP + NTD
For CurveIx:
Cost = 2Topen + mTsel + NfTP + NfTD
If CurveIx is to be worthwhile, we need:
(1) mTsel + NfTP < NPTP
(2) NfTD < NTD
(2) simply requires f < 1, which is essential anyway.
... CurveIx vs. Linear Scan
If we assume that each curve lookup involves 3-4 page access
(i.e. Tsel=3), we can restate (1) as:
(1) (3m+Nf)TP < NPTP
Some observations, given NP = 0.05 N:
- we can't afford to have too many curves (m)
- CurveIx has to do effective filtering (f)
- clustering can reduce the NfTP term
Difficult to analyse further, so we implemented the system to see
how it performed ...
Implementation
Aim: build a quick/easy/light implementation of the above approach.
-
make it stand-alone from QBIC, but use QBIC's distance function
-
use the Unix dbm libraries to implement keyed retrieval
Two components:
build
- reads image idents and color histograms from stdin
- stores each in the Db,
a dbm B-tree keyed on ident
- computes m curve-points for each,
and stores in Index,
a dbm B-tree keyed on curveId
and position (Ci(v) value)
query
- reads a single color histogram from stdin
- uses above method to find k best matches
- display list of best matches idents and distances on stdout,
along with performance statistics
Example for build Program
Inserted img00000
Inserted img00001
...
where the data file looks like:
...
img00102
54 0 0 0 0 0 0 5 10 3 1 1 0 0 0 0
0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0
0 0 0 1 0 0 1 0 0 0 0 0 0 0 0 0
0 1 8 10 3 2 1 7 3 2 1 1 0 0 0 0
0 0 0 2 14 34 3 1 2 1 1 0 0 0 0 0
0 0 0 0 0 0 2 1 0 0 0 3 1 1 2 737
3 0 5 2 1 0 1 0 0 0 0 0 0 0 0 0
0 0 0 0 7 0 25 2 1 1 1 0 0 0 9 2
1 1 0 0 1 2 0 0 0 0 0 0 0 0 0 0
0 0 0 0 0 0 3 0 0 0 5 0 0 0 1 0
0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0
0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0
0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0
0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0
0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0
0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0
...
Example for query Program
An example of executing query on a database of 20,000
images (color histograms) with 100 curves:
QUERY: img00102
0.000000 img00102
0.005571 img00713
0.008826 img00268
0.011413 img05054
0.011811 img00631
0.014259 img04042
0.027598 img00203
0.037471 img00287
0.063639 img00244
0.067583 img00306
# dist calculations: 655
...
various dbm, i/o statistics
...
2.00user 0.27system ...
Ci Values as Keys
One problem raised early in the implementation:
-
the Hilbert numbers produced by Ci
could be expanded to arbitrary precision in order
to distinguish between
Ci(p1 )
and
Ci(p2 )
To use such values as dbm keys, we
need them as fixed precision, so we limit
expansion (to 4 levels).
Problems:
-
gives keys that are 96 bytes long
(producing very large Index files)
-
different vj can be mapped to same point
(size of candidate sets is artificially inflated)
Solution:
-
use a "prefix" index structure (something like a trie)
Query Algorithm
Given: Db , Index , q , k , Range , Ncurves
dbm.open(Db)
dbm.open(Index)
compute Ci perms and shifts
for i in 1..Ncurves
{
p = Ci(vq)
up = dbm.lookup(Index , (i,p))
dn = dbm.lookup(Index , (i,p))
img = dbm.fetch(up)
insert img in results
while within Range for up and dn
{
dbm.next(up); img = dbm.fetch(up)
check insert img in results
dbm.prev(dn); img = dbm.fetch(dn)
check insert img in results
}
}
for i in 1..k
display (dist,imageId) for resultsi
... Query Algorithm
check insert img in results
{
if already seen img
return // no need to check
data = dbm.lookup(Db , img)
vimg = dbm.fetch(data)
d = D(vimg , vq)
include img in k-NN if d small enough
}
The dbm.lookup steps are expensive (Tsel)
Once a cursor is established, access is via cached buffer.
Performance Analysis
Since CurveIx does not guarantee to deliver the k-NN
among its candidates, we set an "acceptable" accuracy level of 90%.
In other words, CurveIx must deliver 0.9 k nearest-neighbours
to be considered useful.
Our initial concern has been to answer the questions:
-
How many curves are needed to achieve 90% accuracy?
-
How many curve-neighbours do we need to examine?
-
Can all of this be done reasonably efficiently?
So far, preliminary answers are only avaliable for 256-d
QBIC color histograms:
- around 80-100
- around 15-30
- not particularly ... yet
Experiments
Measures for accuracy:
- Acc1 average top-10 entries in CurveIx top-10
- Acc2 how frequently CurveIx gives 10 out of 10
Measures for efficiency:
- Size size of Db file + Index file
- Dist number of distance calculations required
- IO total amount of I/O performed
To determine how these measures vary:
- built databases of size 5K, 10K, 15K, 20K (supersets)
- for each database, ran 25 query "benchmark" set
- for each query, ran for 3,5,10,20,30,40 curve-neighbours
(but because of curve-mapping problem, only got 20,30,40)
- for each query, ran for 20,40,60,80,100 curves
Also implemented a linear scan version for comparison and
to collect the exact answer sets.
Sample Comparison
| CurveIx Indexing |
Linear Scan |
QUERY: img00102
0.000000 img00102
0.005571 img00713
0.008826 img00268
0.011413 img05054
0.011811 img00631
0.014259 img04042
0.027598 img00203
0.037471 img00287
0.063639 img00244
0.067583 img00306
# dist calcs: 524
1.67user 0.23system ...
|
QUERY: img00102
0.000000 img00102
0.005571 img00713
0.008826 img00268
0.011413 img05054
0.011811 img00631
0.014259 img04042
0.027598 img00203
0.037471 img00287
0.063639 img00244
0.067583 img00306
# dist calcs: 20000
1.93user 1.12system ...
|
Experimental Results
For fixed database (20K), effect of varying Range, Ncurves
| Curves | Range | Acc1 | Acc2 | Dist |
| 20 | 20 | 6.72 | 0.20 | 426 |
| 20 | 30 | 7.28 | 0.28 | 695 |
| 20 | 40 | 7.68 | 0.36 | 874 |
| 40 | 20 | 7.68 | 0.28 | 957 |
| 40 | 30 | 8.16 | 0.40 | 1301 |
| 40 | 40 | 8.60 | 0.44 | 1703 |
| 60 | 30 | 8.40 | 0.44 | 1905 |
| 60 | 40 | 8.60 | 0.48 | 2413 |
| 80 | 30 | 8.87 | 0.58 | 2485 |
| 80 | 40 | 9.20 | 0.72 | 3381 |
| 100 | 30 | 9.10 | 0.70 | 3061 |
| 100 | 40 | 9.28 | 0.72 | 4156 |
Results: Size vs. Accuracy
For fixed CurveIx parameters (80 curves, 30 range), effect of varying
database size:
| #Images | Acc1 | Acc2 |
| 5K | 9.72 | 0.76 |
| 10K | 9.44 | 0.80 |
| 15K | 9.16 | 0.70 |
| 20K | 9.04 | 0.64 |
Results: Costs
For the 20K database (CurveIx (80,30)):
| Approach | Size(Db) | Size(Ix) | IO(bytes) |
| Linear Scan | 42MB | - | 42MB |
| CurveIx | 46MB | 345MB | 4+36MB |
Other Optimisations for QBIC Indexing
The following techniques can be applied regardless of
the underlying index structure.
They involve exploiting properties of data structures
and access patterns to reduce amount of I/O.
Derived primarily from profiling studies of QBIC.
Caching
Maintain an in-memory collection of relevant pages.
Read/write pages only when absolutely necessary.
Most commercial DBMSs do this invisibly and effectively.
The dbm system:
- performs useful caching while a database file is open
- throws away cache each time file is closed
Current implementation of QBIC opens/closes file for each
record look-up.
... Caching
Interpolation of open-file cache significantly reduces calls
to operating system for file I/O.
Example: insertion of 1000 colour histograms:
...
<1000> Insert of ... successful
Reads: 37504864/13137 Writes: 9383936/2291
1213.8u 49.2s 26:02 80% 1760+4256k 0+0io 2087pf+0w
...
<1000> Insert of ... successful
Reads: 4096/1 Writes: 618496/151
1207.2u 11.8s 24:33 82% 2895+4440k 0+0io 2116pf+0w
Since Unix performs its own caching, this technique
yields little impact on wall-clock times.
Since queries are handled by sequential scan, cache
provides little benefit for them.
For a query server, may provide some querying benefit.
Compression
QBIC colour histograms:
- consist of 256 [0..1000] values as
short[256]
- contain many zero values (on average >200/256)
Scope for compression
- 16-bit quantities used for 10-bit values
(fixed-length compressed vectors)
- because of repetition of particular value (zero)
(variable-length compressed vectors)
Have implemented second:
- reduced storage costs and query I/O by around 2/3
- no increase in processing cost (slight reduction)
Clustering
Reducing to k candidates is no better than
kNO candidates
if each one comes from a separate page
(recall than NO may be larger than 100)
This situation becomes worse when
vectors are smaller,
and so NO becomes larger
Approach: try to ensure that "similar" objects are
placed in the same database page.
Plan to look at how effectively clustering can be performed
for very large numbers of dimensions.
Conclusions
The CurveIx approach shows some potential as a solution for the
HDkNN search problem.
The following aspects require further research and implementation
before this approach could be considered a real solution for k-NN:
-
control of search over curves and neighbours
-
better data structure for curve index
-
clustering strategies to reduce page reads
-
compare to other approaches (e.g.
[Pyramid technique]
)
References/Further Reading
-
Berchtold & Böhm & Kriegel,
The Pyramid Technique: Towards breaking the curse of dimensionality,
ACM SIGMOD, 1998
[proposes a new data access technique which is optimized for high-dimensional
range queries]
-
Gaede & Günther,
Multidimensional access methods,
Technical Report, Humboldt University, Berlin, 1995
[comprehensive survey of multidimensional database access methods]
-
Hellerstein, Koutsoupias & Papadimitriou,
On the analysis of indexing schemes,
ACM PODS, 1997
[initial work on a general theory of "indexability", which is
analogous to complexity theory and describes how effective
data/query/access-method combinations are in terms
of disk access and storage cost]
-
Kurniawati, Jin & Shepherd,
Techniques for supporting efficient content-based retrieval
in multimedia databases,
Australian Computer Journal, 1997
-
Megiddo & Shaft,
Efficient nearest neighbour indexing based on a collection
of space-filling curves,
IBM Internal Research Report, 1997
[description of method to index multidimensional data based on
a multiple complementary "linearisations" of the data points]
-
Weber & Blott,
An Approximation-Based Data Struture for Similarity Search,
Unpublished Report, Bell Labs, 1998
[provides reasonably formal argument that partitioning-based methods
degenerate to worse than linear search once dimensionality around
6..10; proposes VA-files as an alternative]
-
White & Jain,
Algorithms and Strategies for Similarity Retrieval,
UCSD Visual Computing Lab Report, 1996
[survey and characterisation of a large number of approaches
to multidimensional access methods]
Plus numerous other papers downloaded into /home/jas/papers.
Produced: 28 Aug 98
![[Diagram:pic/qbic]](pic/qbic.gif)
![[Diagram:pic/qbic]](pic/qbic.gif)
![[Diagram:pic/cbir-sys]](pic/cbir-sys.gif)
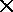 N table of
pre-computed distances to speed up retrieval within database.
N table of
pre-computed distances to speed up retrieval within database.
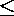 D(va2,vq)
D(va2,vq) 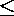 ...
... 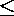 D(vak,vq)
D(vak,vq)
 P/O
P/O
 N/NO
N/NO
![[Diagram:pic/space]](pic/space.gif)
![[Diagram:pic/tree]](pic/tree.gif)
![[Diagram:pic/qspace]](pic/qspace.gif)
![[Diagram:pic/sigfile]](pic/sigfile.gif)
![[Diagram:pic/va-partition]](pic/va-partition.gif)
![[Diagram:pic/c-curves]](pic/c-curves.gif)
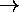 [0,1)
[0,1)
![[Diagram:pic/curves]](pic/curves.gif)