Laboratory
Objectives
- to implement a variant of path finding in a weighted graph
- to see how graphs might be used with real-world data
- to implement a Web crawler
- to build a directed graph structure
Assessment
- Deadline
- 13 January 2019, 23:59:59
- Marks
- 3
- Submission
give cs2521 lab05
Setting up
Create a directory for this lab, change into it,
and retrieve the files with the fetch command:
2521 fetch lab05
This will have provided you with several files (as you may have guessed by its output). They’re described in slightly more detail in the relevant exercises below. Download all the files.
The provided code contains tags marking unused variables. You should remove these tags as you implement those functions.
Exercise 1: Path-Finding (2 marks)
Geographic data is widely available, thanks to sites such as GeoNames. For this lab, we have downloaded data files from the City Distance Dataset by John Burkardt in the Department of Scientific Computing at Florida State University. The dataset that we will use contains information about distances between 30 prominent cities around the world. It measures “great circle” distances: we’ll assume that these measure the distances that an aircraft might fly between the two cities. The data that we have forms a complete graph, in that there is a distance recorded for every pair of cities.
The following diagram shows a subset of the data from the City Distance Dataset.
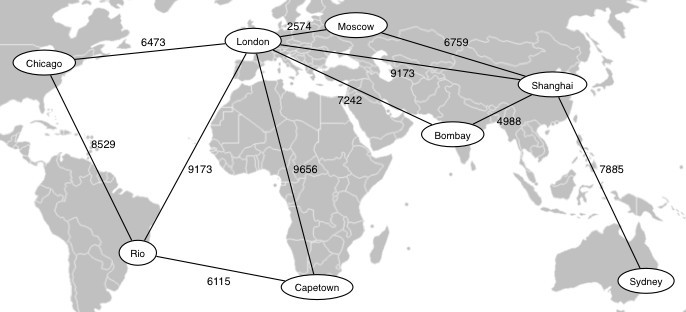
Map from "BlankMap-World-v2" by original uploader: Roke.
Own work. Licensed under Creative Commons Attribution-Share Alike 3.0, via Wikimedia Commons
The data comes in two files:
ha30_dist.txt- This file contains a matrix of distances between cities.
This is essentially the adjacency matrix for a weighted graph
where the vertices are cities and the edge weights correspond
to distances between cities.
As you would expect for an adjacency matrix, the leading diagonal contains all zeroes (in this case, corresponding to the fact that a city is zero distance from itself).
ha30_name.txt- This file contains one city name per line. If we number the lines starting from zero, then the line number corresponds to the vertex number for the city on that line. For example, the Azores is on line 0, so vertex 0 corresponds to the Azores, and the first line in the distance file gives distances from the Azores to the other 29 cities. The last line (line 29) tells us that Tokyo is vertex 29, and the last line in the distance file gives distances between Tokyo and all other cities.
If you’ve done the setup correctly, you should find these files:
Makefile- a set of dependencies used to control compilation
travel.c- main program to load and manipulate the graph
Graph.h- interface to Graph ADT
Graph.c- implementation of Graph ADT
Queue.h- interface to Queue ADT
Queue.c- implementation of Queue ADT
Item.h- definition of Items (Edges)
ha30_name.txt- the city name file described above
ha30_dist.txt- the distance matrix file described above
The Makefile produces a file called travel based on the main program
in travel.c and the functions in Graph.c. The travel program takes
either zero or two command line arguments.
If given zero arguments, it simply displays the graph (in the format described below).
./travel ... displays the entire graph ... ... produces lots of output, so either redirect to a file or use `less' ...
If given two arguments, it treats the first city as a starting point and the second city as a destination, and determines a route between the two cities, based on “hops” between cities with direct flights.
./travel from-city to-city
... display a route to fly between specified cities ...
Read the main() function so that you understand how it works, and,
in particular, how it invokes the functions that you need to implement.
The Graph ADT in this week’s lab implements something that should
resemble a standard adjacency matrix representation of the kind we
looked at in lectures. However, some aspects of it are different to
what we’ve seen in lectures:
Note that city names are not stored as part of the GraphRep data
structure. The Graph ADT deals with vertices using their numeric ID.
The main program maintains the list of city names and passes this list
to the showGraph() function when it is called to display the graph.
This means that the calling interface for the showGraph() function is
different to the showGraph() function from the Graph ADT in
lectures.
The values stored in the matrix are not simply true and false, but represent the distances between vertices. In other words, we’re dealing with a weighted graph. Note, however, that we are not actually using the weights except to indicate the existence of an edge. In this implementation, a weight value of zero indicates no edge between two vertices, while a non-zero weight indicates an edge.
The Graph ADT includes a sub-ADT for Edges. The implementation of
insertEdge() in lectures only required the vertices at the endpoints
of the Edge (i.e. insertEdge(g,v,w)). The version of insertEdge()
for this lab also requires a weight for the edge (i.e.
insertEdge(g,v,w,weight)).
The main program makes some changes to the edges implied by the distance
matrix as it copies them into the Graph. The values in the
ha30_dist.txt file are given in units of “hundreds of miles”; we
want them in units of kilometers so each distance is converted before it
is added to the graph as the weight of an edge.
Since every city has adistance to every other city (except itself),
this gives us a complete graph.
As supplied, the Graph.c file is missing implementations for the
findPath() function. If you compile the travel program and try to
find any route, it will simply tell you that there isn’t one. If you
run travel with no arguments, it will print a representation of the
graph (you can see what this should look like in the file
graph.txt.
You need to implement the findPath(g,src,dest,max,path) function. This
function takes a graph g, two vertex numbers src and dest, a
maximum flight distance, and fills the path array with a sequence of
vertex numbers giving the “shortest” path from src to dest where
no edge on the path is longer than max. The function returns the
number of vertices stored in the path array; if it cannot find a path,
it returns zero. The path array is assumed to have enough space to
hold the longest possible path (which would include all vertices).
This could be solved with a standard BFS graph traversal algorithm, but there are two twists for this application:
-
The edges in the graph represent real distances, but the user of the
travelprogram (the traveller) isn’t necessarily woried about real distances. They are more worried about the number of take-offs and landings (which they find scarey), so the length of a path is measured in terms of the number of edges, not the sum of the edge weights. Thus, the “shortest” path is the one with the minimum number of edges. -
While the traveller isn’t worried about how far a single flight goes, aircraft are affected by this (e.g. they run out of fuel). The
maxparameter forfindPath()allows a user to specify that they only want to consider flights whose length is at mostmaxkilometers (i.e. only edges whose weight is not more thanmax).
Your implementation of findPath() must satisfy both of the above.
In implementing findPath(), you can make use of the Queue ADT that
we’ve supplied. This will create a queue of Vertex numbers.
Note that the default value for max, set in the main program is
10000 km. Making the maximum flight distance smaller produces more
interesting paths (see below), but if you make it too small (e.g.
5000km) you end up isolating Australia from the rest of the world. With
maximum flights of 6000km, the only way out of Australia in this data is
via Guam. If you make the maxmimum flight length large enough (e.g.
aircraft technology improves significantly), every city will be
reachable from every other city in a single hop.
Some example routes (don’t expect them to closely match reality):
# when no max distance is given on the command line, # we assume that planes can fly up to 10000km before refuelling ./travel Berlin Chicago Least-hops route: Berlin ->Chicago ./travel Sydney Chicago Least-hops route: Sydney ->Honolulu ->Chicago ./travel Sydney London Least-hops route: Sydney ->Shanghai ->London ./travel London Sydney Least-hops route: London ->Shanghai ->Sydney ./travel Sydney 'Buenos Aires' Least-hops route: Sydney ->Honolulu ->Chicago ->Buenos Aires # if no plane can fly more than 6000km wihout refuelling ./travel Sydney London 6000 Least-hops route: Sydney ->Guam ->Manila ->Bombay ->Baghdad ->London # if no plane can fly more than 5000km wihout refuelling # you can't leave Australia under this scenario ./travel Sydney 'Buenos Aires' 5000 No route from Sydney to Buenos Aires # if no plane can fly more than 7000km wihout refuelling ./travel Sydney 'Buenos Aires' 7000 Least-hops route: Sydney ->Guam ->Honolulu ->Chicago ->Panama City ->Buenos Aires # planes can fly up to 8000km wihout refuelling ./travel Sydney 'Buenos Aires' 8000 Least-hops route: Sydney ->Guam ->Honolulu ->Mexico City ->Buenos Aires # planes can fly up to 11000km wihout refuelling ./travel Sydney 'Buenos Aires' 11000 Least-hops route: Sydney ->Bombay ->Azores ->Buenos Aires # planes can fly up to 12000km without refuelling # can reach Buenos Aires on a single flight ./travel Sydney 'Buenos Aires' 12000 Least-hops route: Sydney ->Buenos Aires ./travel Bombay Chicago 5000 Least-hops route: Bombay ->Istanbul ->Azores ->Montreal ->Chicago ./travel Sydney Sydney Least-hops route: Sydney
The above routes were generated using an algorithm that checked vertices in order (vertex 0 before vertex 1 before vertex 2, etc.). If you check in a different order, you may generate different, but possibly equally valid, routes.
Make the relevant changes to Graph.c to complete the above tasks.
For this part, you may only submit Graph.c.
Exercise 2: Web-Crawling (1 marks)
We can view the World Wide Web as a massive directed graph, where pages (identified by URLs) are the vertices and hyperlinks (HREFs) are the directed edges. Unlike the graphs we have studied in lectures, there is not a single central representation (e.g. adjacency matrix) for all the edges in the graph of the web; such a data structure would clearly be way too large to store. Instead, the “edges” are embedded in the “vertices”. Despite the unusual representation, the Web is clearly a graph, so the aim of this lab exercise is to build an in-memory graph structure for a very, very small subset of the Web.
Web crawlers are programs that navigate the Web automatically, moving from page to page, processing each page they visit. Crawlers typically use a standard graph traversal algorithm to:
- maintain a list of pages to visit (a ToDo list)
- “visit” the next page by grabbing its HTML content
- scanning the HTML to extract whatever features they are interested in
- collecting hyperlinks from the visited page, and adding these to the ToDo list
- repeating the above steps (until there are no more pages to visit :-)
You need to implement such a crawler, using a collection of supplied
ADTs and a partially complete main program. Your crawler processes each
page by finding any hyperlinks and inserting the implied edges into a
directed Graph ADT based on an adjacency matrix. One difference
between this Graph ADT and the ones we have looked at in lectures is
that you dynamically add information about vertices, as well as edges.
The following diagram shows what an instance of the Graph ADT might
look like:

The Graph data structure allows for maxV vertices (URLs), where
maxV is supplied when graph is created. Initially, there are no
vertices or edges, but as the crawler examines the web, it adds the URLs
of any pages that it visits and records the hyperlinks between them.
This diagram shows what a web crawler might have discovered had it
started crawling from the URL http://example.com/index.html, and so
far examined four web pages.
If we number the four pages from 0..3, with
- page (vertex) 0 being
http://example.com/index.html, - page (vertex) 1 being
http://example.com/category/index.html - page (vertex) 2 being
http://example.com/products.html - page (vertex) 3 being
http://example.com/products/abc.html
The vertices array holds the actual URL strings and also, effectively,
provides the mapping between URLs and vertex numbers. The edges array
is a standard adjacency matrix. The top row tells us that page 0 has
hyperlinks to pages 1 and 2. The second row tells us that page 1 has a
hyperlink back to page 0. The third row similarly shows a hyperlink from
page 2 back to page 0, but also a hyperlink to page 3.
If you’ve done the setup correctly, you should find the following files in your lab directory:
| Makefile | a set of dependencies used to control compilation. |
| crawler.c | main program to crawl and build the graph (not yet complete) |
It also contains .c and .h files for a collection of ADTs: standard
ones such as Set, Stack, Queue and Graph (a directed graph),
along with some ADTs for working with Web data. There are also a set of
files (tg.c, tk.c, tq.c, ts.c) which provide drivers for testing
the various ADTs.
The only file you need to modify is crawler.c, but you need to
understand at least the interfaces to the functions in the various ADTs.
This is described in comments at the start of each function in the .c
files. You can also see examples of using the ADT functions in the
t?.c files. Note that there’s no test file for the HTML parsing and
URL-extracting code, because the supplied version of crawl.c
effectively provides this.
Note that HTML parsing code was borrowed from Dartmouth College. If you looks at the code, it has quite a different style to the rest of the code. This provides an interesting comparison with our code.
The crawl program is used as follows:
./crawl starting-URL max-urls-in-graph
The starting-url tells you which URL to start the crawl from,
and should be of the form http://x.y.z/.
The crawler uses this URL as both the starting point
and uses a normalised version as the base against
which to interpret other URLs.
The max-urls-in-graph specifies
the maximum number of URLs that can be stored in the Graph.
Once this many URLs have been scanned,
the crawling will stop,
or will stop earlier if there are
no more URLs left in the ToDo list.
If you compile then run the supplied crawler on the UNSW handbook, you might see something like:
./crawl http://www.handbook.unsw.edu.au/2017/ 100 Found: 'http://www.unsw.edu.au' Found: 'https://my.unsw.edu.au/' Found: 'https://student.unsw.edu.au/' Found: 'http://www.futurestudents.unsw.edu.au/' Found: 'http://timetable.unsw.edu.au/' Found: 'https://student.unsw.edu.au/node/62' Found: 'http://www.library.unsw.edu.au/' Found: 'http://www.handbook.unsw.edu.au/2017/' Found: 'http://www.unsw.edu.au/faculties' Found: 'https://student.unsw.edu.au/node/4431' Found: 'https://student.unsw.edu.au/node/128' Found: 'http://www.unsw.edu.au/contacts' Found: 'http://www.handbook.unsw.edu.au/general/current/SSAPO/previousEditions.html' Found: 'http://www.handbook.unsw.edu.au/undergraduate/2017/' Found: 'http://www.handbook.unsw.edu.au/postgraduate/2017/' Found: 'http://www.handbook.unsw.edu.au/research/2017/' Found: 'http://www.handbook.unsw.edu.au/nonaward/2017/' Found: 'http://www.unsw.edu.au/future-students/domestic-undergraduate' Found: 'http://www.unsw.edu.au/future-students/postgraduate-coursework' Found: 'http://research.unsw.edu.au/future-students' Found: 'http://www.international.unsw.edu.au/#1' Found: 'https://student.unsw.edu.au/node/1334' Found: 'https://moodle.telt.unsw.edu.au/login/index.php' Found: 'https://student.unsw.edu.au/node/943' Found: 'https://apply.unsw.edu.au/' Found: 'https://student.unsw.edu.au/node/5450' Found: 'http://cgi.cse.unsw.edu.au/~nss/feest/' Found: 'http://www.unsw.edu.au/privacy' Found: 'http://www.unsw.edu.au/copyright-disclaimer' Found: 'http://www.unsw.edu.au/accessibility'
The supplied crawler simply scans the URL given on the command line,
prints out any URLs that it finds, and then stops. It is not attempting
to traverse any further than the supplied page. The second command-line
argument, which limits the size of the Graph is effecitvely ignored
here, since the supplied code does not build a Graph; you need to add
the code to do this.
If you run the supplied “crawler” on http://www.cse.unsw.edu.au, you
don’t get very much, because the CSE website recently moved under the
Engineering Faculty system and the above URL is just a redirection page
to the new site. Copying/pasting the redirection URL gives you more
interesting results.
Before you go running the “crawler” on other websites … DON’T! See the comments below.
HTML is a difficult language to parse, given the way it is frequently (ab)used, and GetNextURL() makes some disgusting approximations which are OK for this lab, but wouldn’t be reasonable in a real Web crawler.
Your task is to modify crawl.c so that it follows any URLs it finds
and builds a Graph of the small region of the Web that it traverses
before bumping in to the max-urls-in-graph limit.
Important: running crawlers outside the UNSW domain is problematic. Running crawlers that make very frequent URL requests is problematic. So:
- DO NOT run your crawler on any website outside UNSW
- YOU MUST include a delay (
sleep(1)) between each URL access
If you don’t do the above, there’s a chance that angry sites that are being hammered by your crawler will block UNSW from future access to those sites. Breaches of the above will result in disciplinary action.
Your crawler can do either a breadth-first or depth-first search, and should follow roughly the graph traversal strategy described in lectures and tutes:
add firstURL to the ToDo list
initialise Graph to hold up to maxURLs
initialise set of Seen URLs
while (ToDo list not empty and Graph not filled) {
grab Next URL from ToDo list
if (not allowed) continue
foreach line in the opened URL {
foreach URL on that line {
if (Graph not filled or both URLs in Graph)
add an edge from Next to this URL
if (this URL not Seen already) {
add it to the Seen set
add it to the ToDo list
}
}
}
close the opened URL
sleep(1)
}
This does not give all the details:
you need to work them out yourself,
based on the supplied ADT functions,
and your understanding of graph traversal.
If you use a Stack for the ToDo list, you’ll do depth-first search.
If you use a Queue for the ToDo list, you’ll do breadth-first search.
A couple more things to note:
(not allowed)refers to not using URLs outside UNSW- the
ToDo listis aStackorQueuerather than a List - if you don’t include the
sleep(1)you will smash whatever web server you access
If you test the crawler out on www.cse.unsw.edu.au, you don’t get
particularly interesting results, because you’ll build a large
adjacency matrix, most of which is empty, before you bump into
MaxURLsInGraph. To assist in doing some feasible crawling and getting
some more interesting output, I have set up a tiny set of self-contained
web pages that you can crawl, starting from:
./crawl http://www.cse.unsw.edu.au/~cs2521/mini-web/ 30
You should use the output of showGraph() to check whether you are
building a plausible Graph. Note that showGraph() has two modes:
showGraph(g,0)shows the URL for each vertex, followed by an indented list of connected verticesshowGraph(g,1)shows just the adjacency matrix in a very compact form; it does not show the stored URLs
Exercise 2: Challenges
There are several aspects of the crawler that you could look to improve:
-
The existing crawler grabs all sorts of URLs that do not represent Web pages. Modify the code so that it filters out non-HTML output.
-
As noted above,
GetNextURL()is a terrible kludge. Modify the code to parse HTML sensibly. -
Modify
showGraph()so that it can (also) produce output that could be fed into a graph drawing tool such as GraphViz or sigmajs, to produce beautiful graph diagrams.
Submitting
You must get the lab marked by your tutor in your lab.
Submit your work with the give command, like so:
give cs2521 lab05 Graph.c crawl.c
Or, if you are working from home, upload the relevant file(s) to the lab05 activity on WebCMS3 or on Give Online.